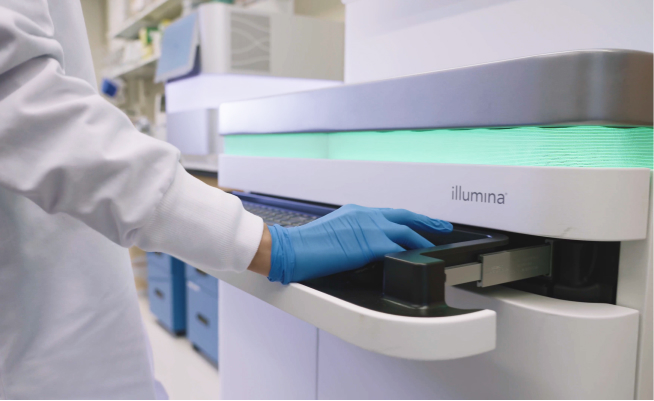
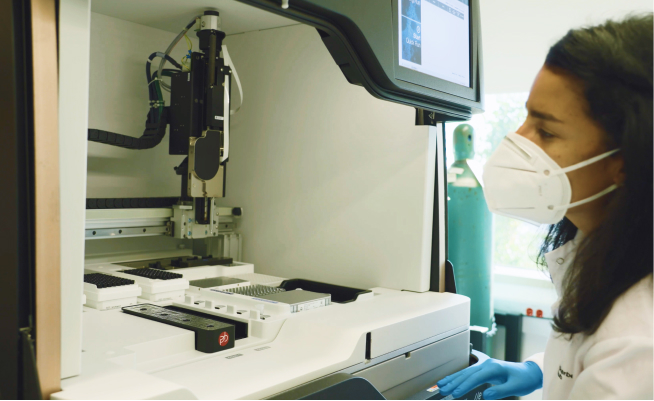
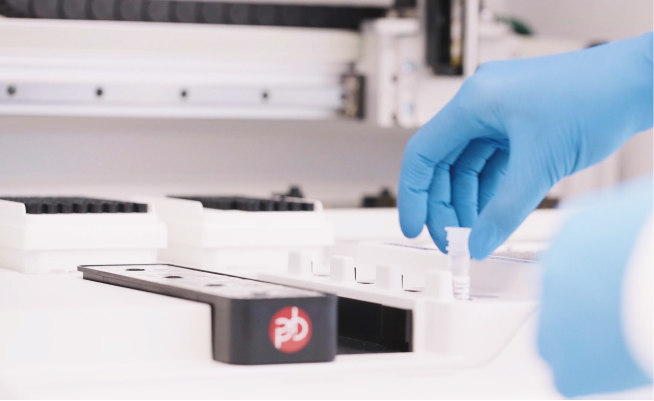
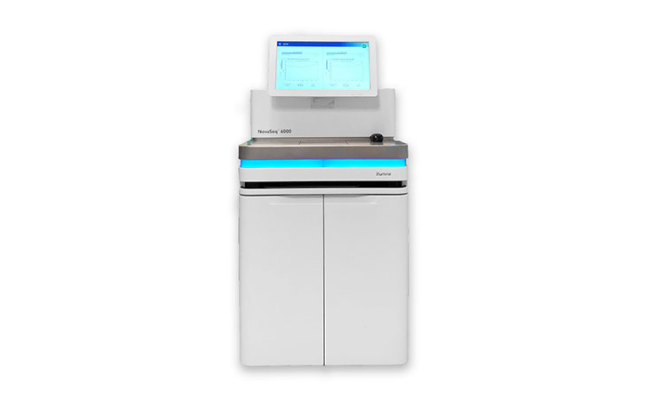
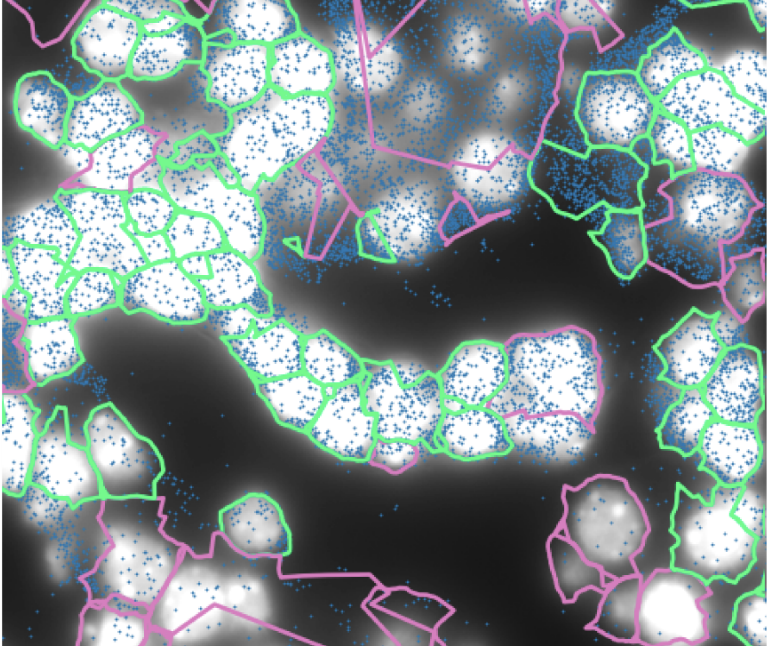
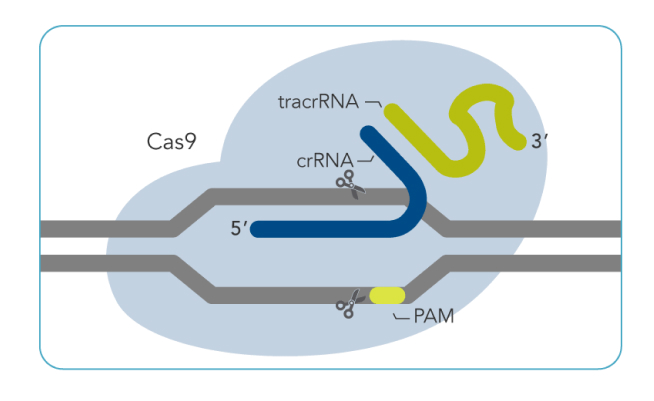
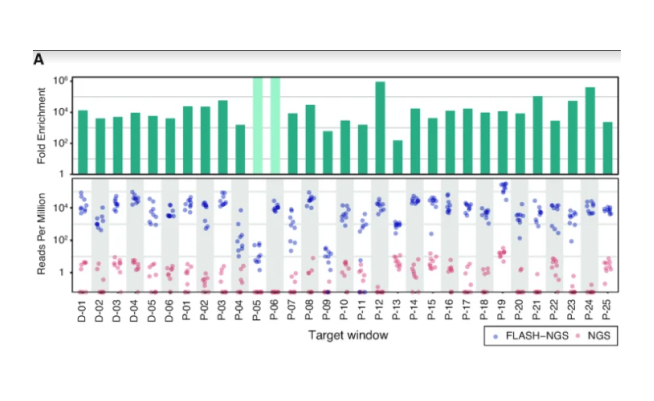
shrimPy is a pythonic framework for correlative 3D imaging of physical and molecular properties of cells at high throughput.

Tools
In the CZ Biohub Network, we know that open access to tools, data, and other resources expedites solutions. Not only do we invent tools to enable avenues for biomedical research that were not possible before, but we also make them available for the scientific community to access and use. We also collaborate closely with the scientific community to support open source tools that are solving the unique analysis needs of researchers.

Rapid QC-MS is an all-in-one solution for automated quality control of liquid chromatography-mass spectrometry instrument runs, both during and after data acquisition.
GET BIOHUB UPDATES IN YOUR INBOX
Stay up-to-date on the latest news, publications, competitions, and stories from CZ Biohub.