Our Research
Led by Norma Neff, the Genomics Platform develops and deploys state-of-the-art protocols in single-cell sequencing with Illumina, 10X, Oxford Nanopore, and PacBio platforms to generate high-quality datasets for cell-type identification and functional analyses as well as for metagenomic, infectious disease, and space biology. Various other emerging technologies such as spatial transcriptomics are being developed and optimized.
Research Projects
Incorporating Emerging Sequencing Technologies
 DNA sequencing is a key measurement technique for many projects at CZ Biohub SF. One research objective of the Genomics Platform is to continuously evaluate and integrate cutting-edge sequencing methodologies to enable utilization of high-quality genomic measurements in new areas of biomedical research. Examples include optimized pipelines incorporating long read technologies that have resulted in strain-level identification of SARS-CoV-2, closed genomes of novel organisms, and produced high-quality profiles of the human immune repertoire. Other examples include spatial sequencing and emerging sequencing instruments.
DNA sequencing is a key measurement technique for many projects at CZ Biohub SF. One research objective of the Genomics Platform is to continuously evaluate and integrate cutting-edge sequencing methodologies to enable utilization of high-quality genomic measurements in new areas of biomedical research. Examples include optimized pipelines incorporating long read technologies that have resulted in strain-level identification of SARS-CoV-2, closed genomes of novel organisms, and produced high-quality profiles of the human immune repertoire. Other examples include spatial sequencing and emerging sequencing instruments.
Single Cell Multi-Omics
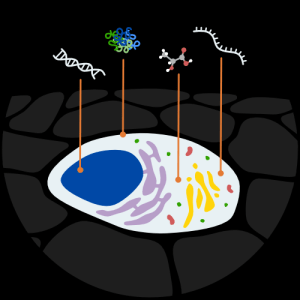 Single-cell research is a core focus at CZ Biohub. The Genomics Platform maintains and continues to improve all droplet- and plate-based single-cell RNA-seq pipelines, enabling their use on a greater variety of organisms and cell types. Projects that benefited from improved sample preservation and dissociation processes developed by the team include the identification of progenitors in zebrafish, delineation of bacterial induced changes in host immune transcription, and atlases of model and non-model organisms.
Single-cell research is a core focus at CZ Biohub. The Genomics Platform maintains and continues to improve all droplet- and plate-based single-cell RNA-seq pipelines, enabling their use on a greater variety of organisms and cell types. Projects that benefited from improved sample preservation and dissociation processes developed by the team include the identification of progenitors in zebrafish, delineation of bacterial induced changes in host immune transcription, and atlases of model and non-model organisms.
Synthetic Gut Microbiota
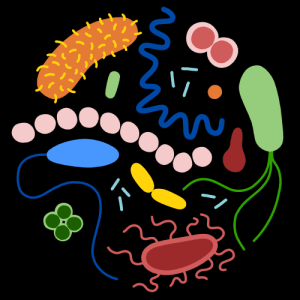 Synthetic microbial communities constructed using pure human gut strains are popular vehicles to explore species interactions and to develop defined synthetic gut communities as potential therapeutics. Genomics Platform members have developed high-throughput pipelines to construct, cultivate, and measure genomes, transcriptomes, and metabolomes of complex but defined communities to study rules of community assembly and resource utilization. The team also builds computational tools with strain-level accuracy based on databases of 100+ reference strains whose genomes have been closed with various sequencing capabilities.
Synthetic microbial communities constructed using pure human gut strains are popular vehicles to explore species interactions and to develop defined synthetic gut communities as potential therapeutics. Genomics Platform members have developed high-throughput pipelines to construct, cultivate, and measure genomes, transcriptomes, and metabolomes of complex but defined communities to study rules of community assembly and resource utilization. The team also builds computational tools with strain-level accuracy based on databases of 100+ reference strains whose genomes have been closed with various sequencing capabilities.
Platform Collaborations
Tabula Projects
Creating transcriptomic maps (atlases) of model organisms at the single-cell level is a central mission of CZ Biohub SF. Members of the Genomics Platform carry out data generation activities for all single-cell Tabula projects (Tabula Muris, Tabula Muris Senis, Tabula Drosophilae, Tabula Danio Rerio [zebrafish], Tabula Sapiens, and a human COVID atlas). In addition, the Genomics Platform facilitates data sharing and supports the construction of data exploration portals.
Infectious Disease
Rapidly identifying and tracking infectious diseases locally and around the world is another central mission of CZ Biohub SF. Members of the Genomics Platform support the metadata tracking and data acquisition efforts of various disease surveillance efforts. In addition, as part of the joint initiative with the Bill & Melinda Gates Foundation, team members train visiting scientists from around the world on best practices of carrying out sequencing projects.
Sequencing Operations
The Genomics Platform currently maintains a lineup of DNA sequencers at Biohub San Francisco. The optimized sequencing workflows and technological capabilities are accessible to internal groups and selected external investigator projects via the Biohub sequencing submission infrastructure.


